Prof. creates first-ever 3-D image of Alzheimer's protein clump
Biophysicist Nikolaus Grigorieff an expert on high-resolution electron microscopy
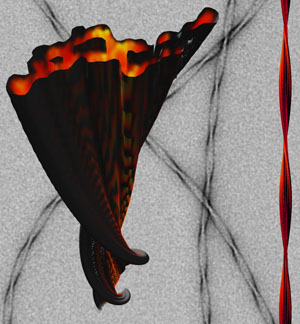 In an important step toward demystifying the role protein clumps play in the development of neurodegenerative disease, researchers have created a stunning three-dimensional picture of an Alzheimer’s peptide aggregate using electron microscopy. The study, in this week’s issue of the Proceedings of the National Academy of Sciences, reports that researchers from Brandeis University in Waltham, Mass., and the Leibniz Institut in Jena, Germany, have shown—for the first time—how A-beta peptide, found in the brains of Alzheimer’s patients, forms a spaghetti-like protein mass called an amyloid fibril.
In an important step toward demystifying the role protein clumps play in the development of neurodegenerative disease, researchers have created a stunning three-dimensional picture of an Alzheimer’s peptide aggregate using electron microscopy. The study, in this week’s issue of the Proceedings of the National Academy of Sciences, reports that researchers from Brandeis University in Waltham, Mass., and the Leibniz Institut in Jena, Germany, have shown—for the first time—how A-beta peptide, found in the brains of Alzheimer’s patients, forms a spaghetti-like protein mass called an amyloid fibril.
Read the UPI story about this research.
“This study is a significant advance regarding our understanding of how these fibrils are built from the A-beta peptide (Alzheimer's peptide),” said co-author Nikolaus Grigorieff, a biophysicist at Brandeis University and an investigator with the Howard Hughes Medical Institute. “People have been guessing for decades what these fibrils look like, but now we have an actual 3D image.”
In healthy people A-beta peptide does not aggregate, but in Alzheimer’s patients it clumps first and then forms long fibrils, like tentacles, in a so-called cross-beta structure. Scientists disagree whether it is the clumps that kill neurons in the brain or the fibrils. Grigorieff wants to discover which part of the amyloid structure is toxic; that would be an important step in designing drugs to prevent or treat disease.
Amyloid structure—the particular way a protein or peptide clumps together—is linked to other neurodegenerative conditions as well, including Parkinson’s and Creutzfeldt-Jakob disease. “The amyloid way of folding and aggregation seems to be a fundamental property of proteins and peptides” explained Grigorieff. “We know how most normal proteins fold, but what drives amyloid formation?”
It’s a question that has dogged structural biologists and biochemists for a long time but stubbornly refuses elucidation. Researchers using x-ray crystallography have so far been unable to obtain crystals from fibrils of full-length polypeptide chains. Structural models based on NMR data have also come up short. Scientists have made do with studying fragments of the A-beta peptide. The major barrier to determining the structure of A-beta fibrils is that the same peptide will exasperatingly assemble differently from fibril to fibril—unlike normal proteins, which reliably fold the same way every time.
Grigorieff and his colleagues overcame this barrier by generating fibrils in a test tube under conditions that reduce the variability between fibrils, and by selecting about 200 images of fibrils that were most similar to each other and averaging them on a computer.
An expert at high-resolution electron cryo-microscopy of protein complexes and macromolecular machines, Grigorieff said his lab made an image of the A-beta fibril at a resolution of eight Angstroms, revealing useful detail at a magnification of roughly a million times. Short of atomic resolution by a factor of 2.5 to 3, the image revealed how the peptide, a series of linked amino acids, was arranged in the tape-like fibril.
“The next step will be a 3D image that tells us exactly where all the amino acids are,” postulates Grigorieff. “This will tell us more about the chemical and biological properties of A-beta fibrils that we need to know to understand their role in Alzheimer’s.”





