Chandler Fulton helps write an amoeba's amazing life story
A one-celled wonder, Naegleria gruberi just filled an important chapter in the book of life
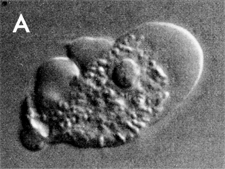 |
| Amoeba |
The single-celled soil amoeba known as Naegleria gruberi, common denizen of mud, puddles, and fresh-water ponds from Brandeis to Bora Bora, Antarctica to the Arctic, is strikingly versatile and evolved for a protozoan. This little eukaryote (organism with a nucleus and other compartmentalized organelles) lives a complex lifestyle that includes walking, swimming, and even turning into a hard protective cyst in unfavorable environments.
In fact, a breakthrough study published as a "Featured Article" in the March 5 issue of Cell by two dozen scientists, including Brandeis Professor of Biology Emeritus Chandler Fulton, reveals that humans and Naegleria have more in common than previously thought. In the study, scientists sequenced the genome of this nifty organism for the first time, raising the curtain on our shared genetic history and the ancient transition from prokaryotes, including bacteria which function with their proteins just floating around in their cells, to more evolved eukaryotes.
 |
|
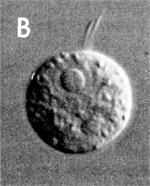
Cell undergoing |
For Fulton, the study’s findings are particularly gratifying, capping an affectionate half-century relationship with these “little guys,” and creating research possibilities for many, many years. In the early sixties, Fulton, an ambitious young biologist at Brandeis, was looking for a model organism in which to study cell differentiation to better understand problems of embryonic development.
“I needed a simple system and I really went searching,” says Fulton. “I found Naegleria gruberi, which hadn’t been studied very much, but as soon I worked with it in the lab I knew it was the perfect organism to use. I was looking for something that could undergo a quick change from one form to a distinctly different form, and this fellow could do that. In an hour and a half it can go from walking around using pseudopods to a streamlined cell with flagella that swims 100 times as fast as amoebae can walk,” says Fulton.
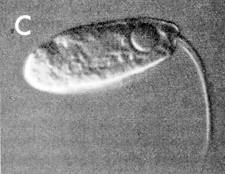 |
| A flagellate |
Over the decades, Fulton and his lab made a number of important discoveries and published about a hundred papers using Naegleria. Among the achievements, the lab developed the standard conditions for growing amoebae and studying their differentiation to flagellates. Fulton and his colleagues discovered that the centriole structures that form at the base of the flagella occur de novo; until then, scientists believed that centrioles could only be produced next to a parent structure.
With the unexpected discovery that the tubulins used to assemble the flagella are synthesized during differentiation and are different from those used for cell division, Fulton proposed the “multitubulin hypothesis,” which influenced a lot of work on the cytoskeleton in the 1980s. All this research established Naegleria as a model system for studying cell differentiation and cell organelle morphogenesis, the process by which cellular components take shape.
In short, Fulton found the amoeba a goldmine for scientific study—a career maker for him, as well as the many undergraduates, graduate students, and postdocs who shared in this research and others who steadfastly studied the protozoan in search of clues about the organization and development of life. For them, the little amoeba has not disappointed.
 |
| Chandler Fulton |
Fast forward 50 years to the publication of the Cell study, which revealed that Naegleria contains almost 16,000 genes coding for proteins, while humans have 23,000 protein-coding genes. Awash in this information, the scientists had to decide which of the thousands of genes were biologically important to explore further. Fulton became involved in the analysis once the genome had been annotated. As an example, he says that the surprising discovery of genes for sexuality is of particular interest, since the organism is only known to reproduce asexually—at a tidy pace of once every hour and a half.
This treasure trove of genes drives many of Naegleria’s other talents, too. For example, the amoeba prefers to lazily lumber about using pseudopods to hunt bacteria. But if walking about (Fulton insists the amoeba does not “ooze”) doesn’t lead to enough food, the amoeba quickly becomes a flagellate. Sporting two whip-like tails that make it the single-celled equivalent of an Olympic swimmer, it can swim off in search of a bigger bacterial buffet before reverting to amoeba-hood to eat and reproduce.
Finally, if conditions become really inhospitable, Naegleria simply hibernates, becoming a walled resting cyst that can endure in unfavorable conditions until dampness, warmth and food bring back the good times and the amoeba becomes its slimy self again. The amoebae can grow “breathing” air, using oxygen for energy as we do, or in the mud at the bottom of a pond, where it has to make energy without oxygen.
The genome was sequenced by the Department of Energy’s Joint Genome Institute, and annotated by a team there and at the University of California, Berkeley, and subsequently analyzed by a team that also included researchers from England, Canada, and Brandeis. The Berkeley researchers compared the Naegleria genome to those of 16 other eukaryotes, ranging from humans and fungi to green plants and other single-celled eukaryotes, revealing for the first time a set of at least 4,000 genes that were probably part of the first, most primitive ancestral eukaryote.
“We know that the lineage of ancestors that led to humans separated from the lineage that produced Naegleria in evolution about 1.5 billion years ago, when eukaryotes were just getting started,” says Fulton. “We went our separate ways, and nobody knows what our common ancestor, the first eukaryote, looked like.”
But comparing the human and amoeba genomes revealed some very surprising findings.
“Naegleria and humans are about as far apart as contemporary eukaryotes can get, but when we compare the genes and find that Naegleria and humans both have the genetic tool kit for mating and reproduction—that tells us that our common ancestor probably had that tool kit, too,” says Fulton, adding, “such deductions mean that those ancient ancestors playing around in low-oxygen slime had many of the same genes that we have today.”
Naegleria’s genes also shed light on how cells move, signal one another, and metabolize nutrients. In fact, Naegleria contains much of the tool kit of genes that distinguish eukaryotes from bacteria, including genes involved in DNA replication and repair, gene expression, intron splicing, metabolic versatility, amoeboid locomotion and flagellate swimming, sexuality, and signaling pathways.
The sequenced genome of Naegleria gruberi opens a new chapter in the book of life, and that manual will guide all future researchers utilizing this organism. Indeed, as the authors wrote in Cell to underscore the paradigm shift the study produced, “it was a giant step to an amoeba, yet a small step to man.”
No wonder Fulton believes—and cartoonist Larson showed—that the amoeba “is very lovable” and not so lowly after all.





