Liz Hedstrom

Professor of Biology and Chemistry
Research Description
Targeted protein degradation, antimicrobial discovery; enzyme structure-function studies, mTOR inhibitors
My laboratory uses approaches derived from both chemistry and biology. Projects include problems in inhibitor design, enzyme catalysis and protein degradation. Techniques vary with the particular project, and can entail molecular biology, organic synthesis, protein crystallography and NMR spectroscopy as well as protein purification, enzyme kinetics and mutagenesis. Our ongoing projects are outlined below. For more information, please see the lab website.
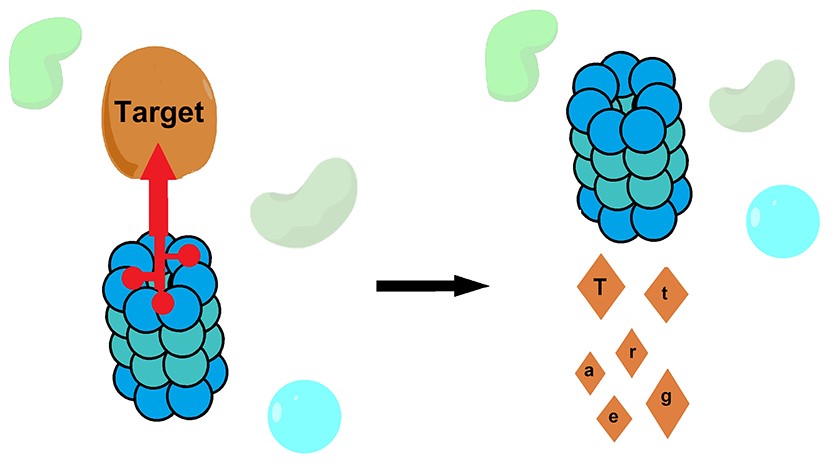 Targeted protein degradation. Targeted protein degradation is an emerging strategy in drug discovery. The rational design of degraders has generally focused on a single strategy: localization of the target protein to a ubiquitin E3 ligase. We have pioneered strategies for ubiquitin-independent degradation, where a small molecule localizes the target protein directly to the proteasome. We expect that this work will also yield important new insights into the pathways of protein quality control.
Targeted protein degradation. Targeted protein degradation is an emerging strategy in drug discovery. The rational design of degraders has generally focused on a single strategy: localization of the target protein to a ubiquitin E3 ligase. We have pioneered strategies for ubiquitin-independent degradation, where a small molecule localizes the target protein directly to the proteasome. We expect that this work will also yield important new insights into the pathways of protein quality control.
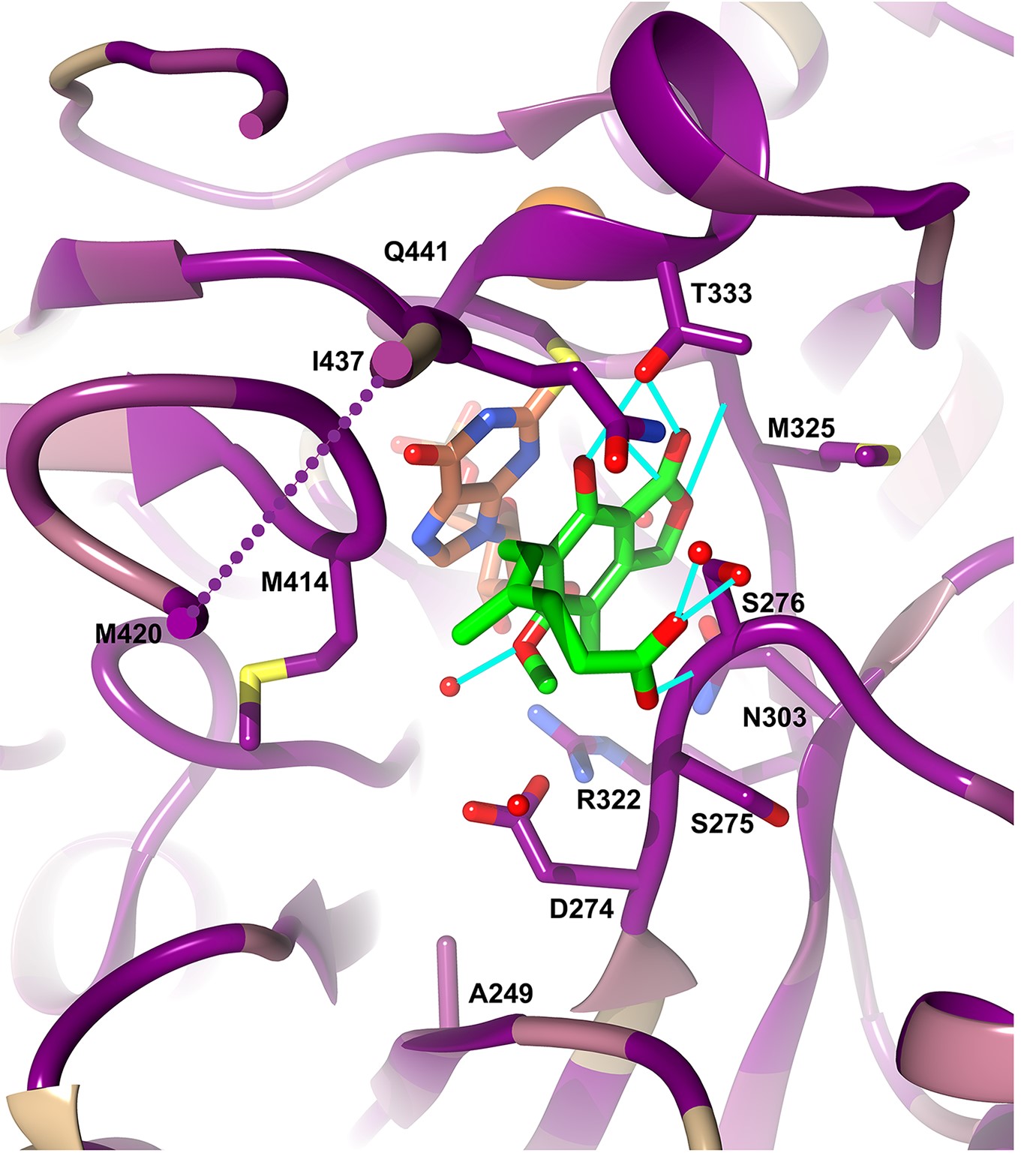 IMPDH-targeted drug discovery for infectious diseases. We exploited the unexpected divergence of the cofactor binding site in IMPDH to identify pathogen-selective inhibitors. Importantly, these compounds also display antibacterial activity. This work could lead to novel treatments for a wide variety of bacterial infections, including some of the most devastating human pathogens such as Mycobacterium tuberculosis. We are also investigating the role of IMPDH in COVID infection in collaboration with the Kadener laboratory at Brandeis. Lastly, we are investigating the mechanism of action of a potent antibacterial compound in collaboration with the Bisson laboratory.
IMPDH-targeted drug discovery for infectious diseases. We exploited the unexpected divergence of the cofactor binding site in IMPDH to identify pathogen-selective inhibitors. Importantly, these compounds also display antibacterial activity. This work could lead to novel treatments for a wide variety of bacterial infections, including some of the most devastating human pathogens such as Mycobacterium tuberculosis. We are also investigating the role of IMPDH in COVID infection in collaboration with the Kadener laboratory at Brandeis. Lastly, we are investigating the mechanism of action of a potent antibacterial compound in collaboration with the Bisson laboratory.
mTOR inhibitors. The protein kinase mTOR is the master regulator of metabolism, growth and longevity. mTOR inhibitors prolong lifespan and are used as immunosuppressive and anticancer agents. We serendipitously discovered a compound that inhibits mTOR signaling by activating the negative regulator TSC2. We are currently unraveling the mechanism of inhibition in collaboration with the Marr laboratory.
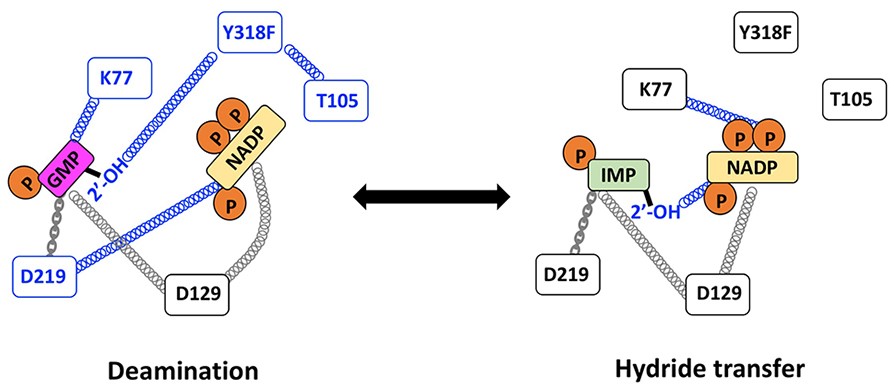 Enzyme evolution. Our current system of interest is the IMPDH/GMPR family, where we are investigating how enzymes with nearly identical active sites produce different reaction outcomes. We are also exploring how antibiotic self-resistance evolves in collaboration with the Theobald laboratory.
Enzyme evolution. Our current system of interest is the IMPDH/GMPR family, where we are investigating how enzymes with nearly identical active sites produce different reaction outcomes. We are also exploring how antibiotic self-resistance evolves in collaboration with the Theobald laboratory.
Selected Publications
- Pepi MJ, Chacko S, Marqus GM, Singh V, Wang Z, Planck K, Cullinane RT, Meka PN, Gollapalli DR, Ioerger TR, Rhee KY, Cuny GD, Boshoff HIM, Hedstrom L. A d-Phenylalanine-Benzoxazole Derivative Reveals the Role of the Essential Enzyme Rv3603c in the Pantothenate Biosynthetic Pathway of Mycobacterium tuberculosis. ACS Infect Dis. 2022 Jan 11. doi: 10.1021/acsinfecdis.1c00461.
- Hedstrom L, Liechti G, Goldberg JB, Gollapalli DR. The antibiotic potential of prokaryotic IMP dehydrogenase inhibitors. Curr Med Chem. 2011;18(13):1909-18.
- Patton GC, Stenmark P, Gollapalli DR, Sevastik R, Kursula P, Flodin S, Schuler H, Swales CT, Eklund H, Himo F, Nordlund P, Hedstrom L. Cofactor mobility determines reaction outcome in the IMPDH and GMPR (beta-alpha)8 barrel enzymes. Nat Chem Biol. 2011;7(12):950-8.
- Sun XE, Hansen BG, Hedstrom L. Kinetically Controlled Drug Resistance: how Penicillium survives mycophenolic acid. J Biol Chem. 2011;286(47):40595-600.
- Gorla SK, Kavitha M, Zhang M, Liu X, Sharling L, Gollapalli DR, Striepen B, Hedstrom L, Cuny GD. Selective and potent urea inhibitors of cryptosporidium parvum inosine 5'-monophosphate dehydrogenase. J Med Chem. 2012;55(17): 7759-71.
- Long MJ, Gollapalli DR, Hedstrom L. Inhibitor mediated protein degradation. Chem Biol. 2012;19(5):629-37.
- Mandapati K, Gorla SK, House AL, McKenney ES, Zhang M, Rao SN, Gollapalli DR, Mann BJ, Goldberg JB, Cuny GD, Glomski IJ, Hedstrom L. (2014). Repurposing cryptosporidium inosine 5'-monophosphate dehydrogenase inhibitors as potential antibacterial agents. ACS Med Chem Lett 5(8): 846-850.
- Makowska-Grzyska M, Kim Y, Maltseva N, Osipiuk J, Gu M, Zhang M, Mandapati K, Gollapalli DR, Gorla SK, Hedstrom L, Joachimiak A. (2015). A Novel Cofactor-binding Mode in Bacterial IMP Dehydrogenases Explains Inhibitor Selectivity. J Biol Chem 290(9): 5893-5911.
- Lawson AP, Long MJ, Coffey RT, Qian Y, Weerapana E, El Oualid F, Hedstrom L. (2015). Naturally Occurring Isothiocyanates Exert Anticancer Effects by Inhibiting Deubiquitinating Enzymes. Cancer Res 75(23): 5130-5142.
- Coffey RT, Shi Y, Long MJ, Marr MT 2nd, Hedstrom L. (2016). Ubiquilin-mediated Small Molecule Inhibition of Mammalian Target of Rapamycin Complex 1 (mTORC1) Signaling. J Biol Chem 291(10): 5221-5233.
- Shi Y, Long MJ, Rosenberg MM, Li S, Kobjack A, Lessans P, Coffey RT and Hedstrom L (2016). Boc3Arg-linked ligands induce degradation by localizing target proteins to the 20S proteasome. ACS Chem Biol. 2016, 11 (12), pp 3328–3337.
- Wei Y, Kuzmic P, Yu R, Modi G and Hedstrom L (2016). Inhibition of Inosine-5'-monophosphate Dehydrogenase from Bacillus anthracis: Mechanism Revealed by Pre-Steady-State Kinetics. Biochemistry 55(37): 5279-5288.
- Chacko S, Boshoff HIM, Singh V, Ferraris DM, Gollapalli DR, Zhang M, Lawson AP, Joachimiak A, Rizzi M, Mizrahi V, Cuny GD and Hedstrom L. Expanding benzoxazole based inosine 5’-monophosphate dehydrogenase (IMPDH) inhibitor structure-activity as potential anti-tuberculosis agents. J. Med. Chem. 61, 4739-4756 (2018).
- Rosenberg MM, Yao T, Patton GC, Redfield AG, Roberts, MF and Hedstrom L. Enzyme-substrate-cofactor dynamical networks revealed by high resolution field cycling relaxometry. Biochemistry 59, 2359-2370 (2020).
- Modi G, Marqus GM, Vippila MR, Gollapalli DR, Kim Y, Manna AC, Shibin C, Maltseva N, Wang X, Cullinane RT, Zhang Y, Kotler JLM, Kuzmic P, Zhang M, Lawson AP, Joachimiak A, Cheung A, Snider BB, Rothstein DM, Cuny GD and Hedstrom L. The enzymatic activity of inosine 5’-monophosphate dehydrogenase may not be a vulnerable target for Staphylococcus aureus infections. ACS Infect Dis. 2021 Nov 12;7(11):3062-3076. doi: 10.1021/acsinfecdis.1c00342
View Complete Publication List on PubMed: Liz Hedstrom