TRIBE: Revolutionizing RNA Target Research
When it comes to developing effective treatments for many cancers and neurological diseases, one barrier has been the speed, efficiency, and accuracy of reliable identification of drug targets. Debilitating disease like myotrophic Lateral Sclerosis (ALS), Fragile X Syndrome (FXS), and Parkinson’s have been have been linked to aberrations in RNA-binding proteins (RBPs). RBPs represent approximately 10% of protein-coding genes in humans, making them of critical importance in the development of new treatments. Identifying RBP-RNA interactions responsible for disease states has been an ongoing challenge for researchers, impeding the path to treatment.
One reason for this is that it’s especially challenging to analyze RBPs with just a small number of cells. The gold-standard method for characterizing RBP–RNA interactions, cross-linking immunoprecipitation (CLIP), requires millions of cells and is vulnerable to false positive signals.
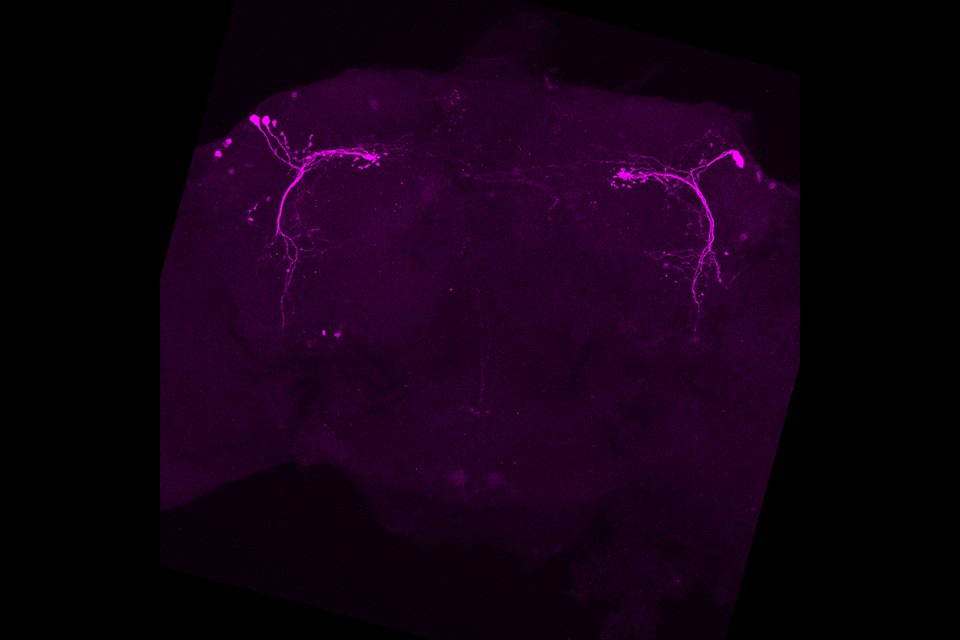
Researchers here at Brandeis in Nobel Prize winner Michael Rosbash’s lab have developed a solution to this conundrum: TRIBE (Targets of RNA-binding proteins Identified By Editing) was designed to solve this problem and it has been proven to be effective even in a small numbers of circadian neurons of the Drosophila adult brain. To date, it has been effective in identifying RBP targets using as few as 150 cells. This is a breakthrough in creating faster and more cost-effective ways of analyzing these proteins, with even greater accuracy, as well. The goal is ultimately to use TRIBE as a means of catalyzing novel discoveries in biology and human disease.
The diverse team behind this innovation is excited to see where it’s heading. “People didn’t think it would work,” remembers Reazur Rahman, PhD, Joint Postdoctoral Research Associate in the Michael Rosbash lab. Along with Weijin Xu, a 5th-year doctoral student, and undergraduate student Josh Lepson, who work together on the project, Reazur notes that “TRIBE can push the needle along” as biotech strives to create new drugs that target RNA molecules, a key emerging area of drug development.
In addition, the team is working on a computational component for HyperTRIBE. HyperTRIBE genetic construct expresses an RBP of interest fused to mutated catalytic domain of the RNA editing enzyme ADAR, which marks the target RNA transcript by editing; these editing events can be identified by sequencing the transcriptome. Currently, both CLIP and HyperTRIBE software identify a set of RNA transcripts that interact with the queried RBP but it is unable to rank these transcripts based on the RBP-RNA binding affinity. HyperTRIBE software
The ranked transcript list from HyperTRIBE will be tremendously beneficial to researchers investigating RBPs role in disease, as it will allow them to focus on selected targets implicated in disease states, catalyzing key advancements in pharmaceutical research.